Overlays#
Overlays generate custom colouring for elements and interactions in the map.
Working with overlays#
Users can view and download available overlays. By pressing “View”, the background of map automatically changes removing any background colour to better emphasise the overlay’s colour. For users that are not logged in, the “Download” button allows you to download the dataset used to generate the overlay. For the logged users who submitted their overlays, the three dot button allows you to edit or download a given overlay.
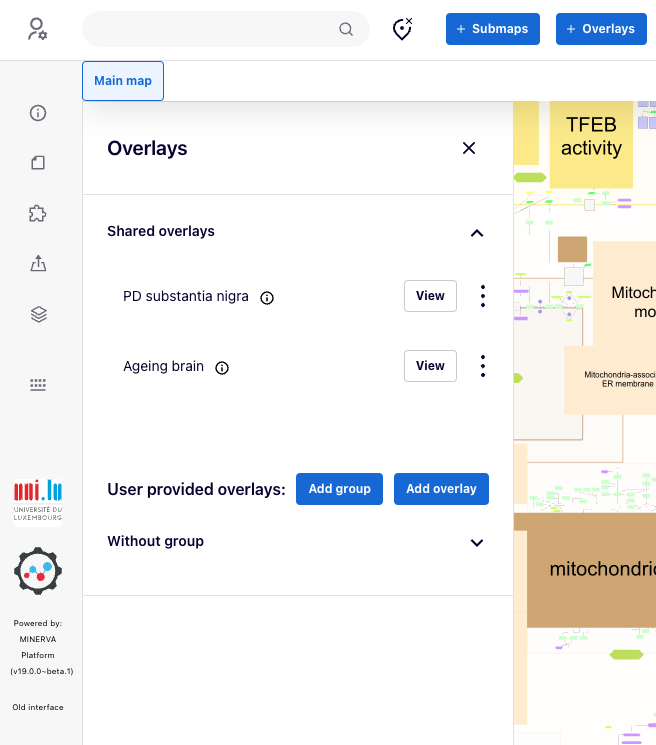
Overlays of custom colours#
Overlays assign fixed colours to elements. Within a single overlay, more than one colour can be assigned to a single element. This multiple colouring will be shown as horizontal stripes. More than one overlay can be viewed at a time, with vertical segments corresponding to each overlay.
After clicking on an element of the map, overlay data for this element are visible in the left panel as one or more horizontal colour bars.
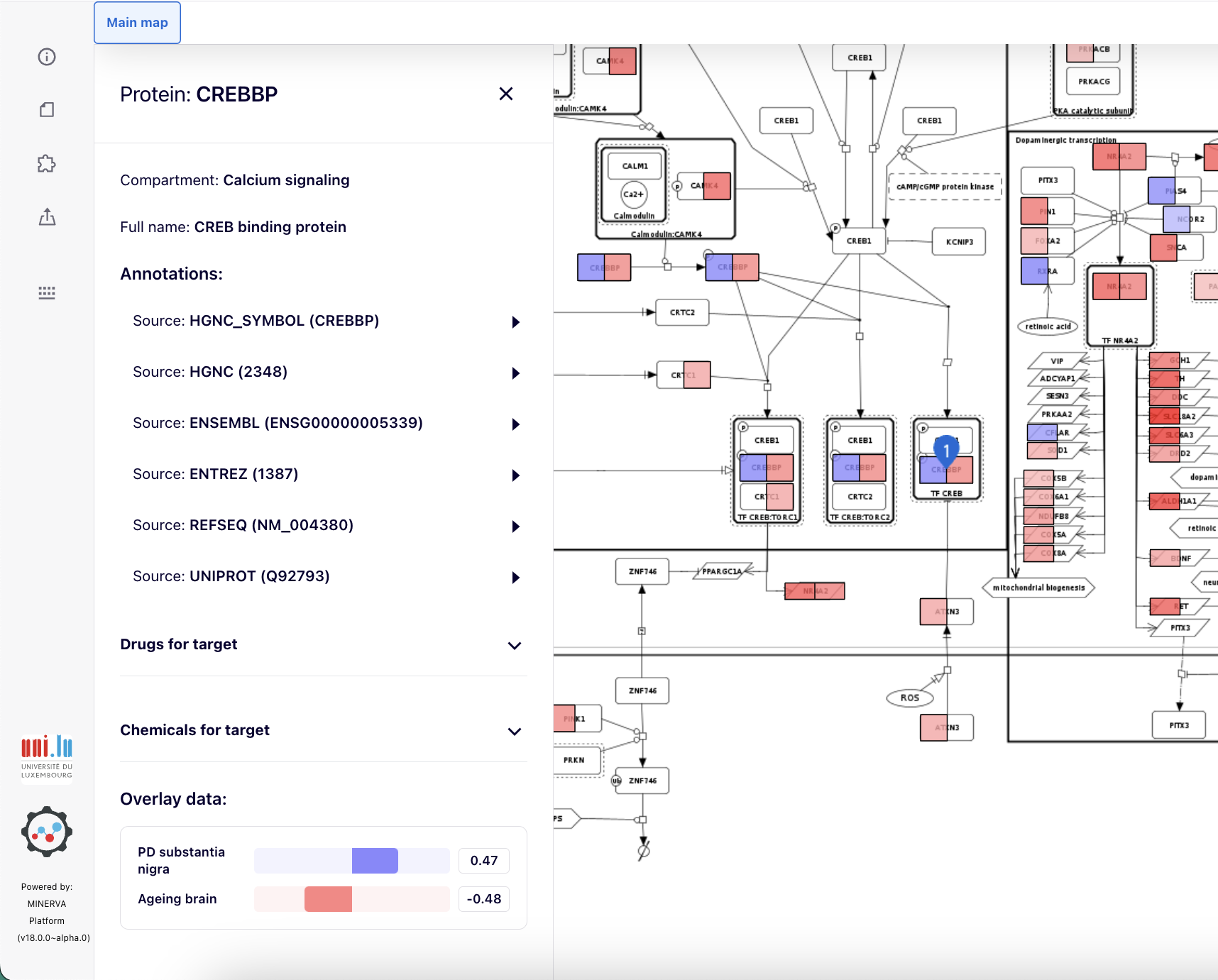
If an element is a submap link, its colouring reflects a summary of the overlay values in the underlying submap. Also, the left panel will display a graphical summary of submap elements and their overlay values Image below shows a data overlay for a group of submap links, each colouring specific to the different submap they link to. Details are shown for the dopaminergic transcription element.
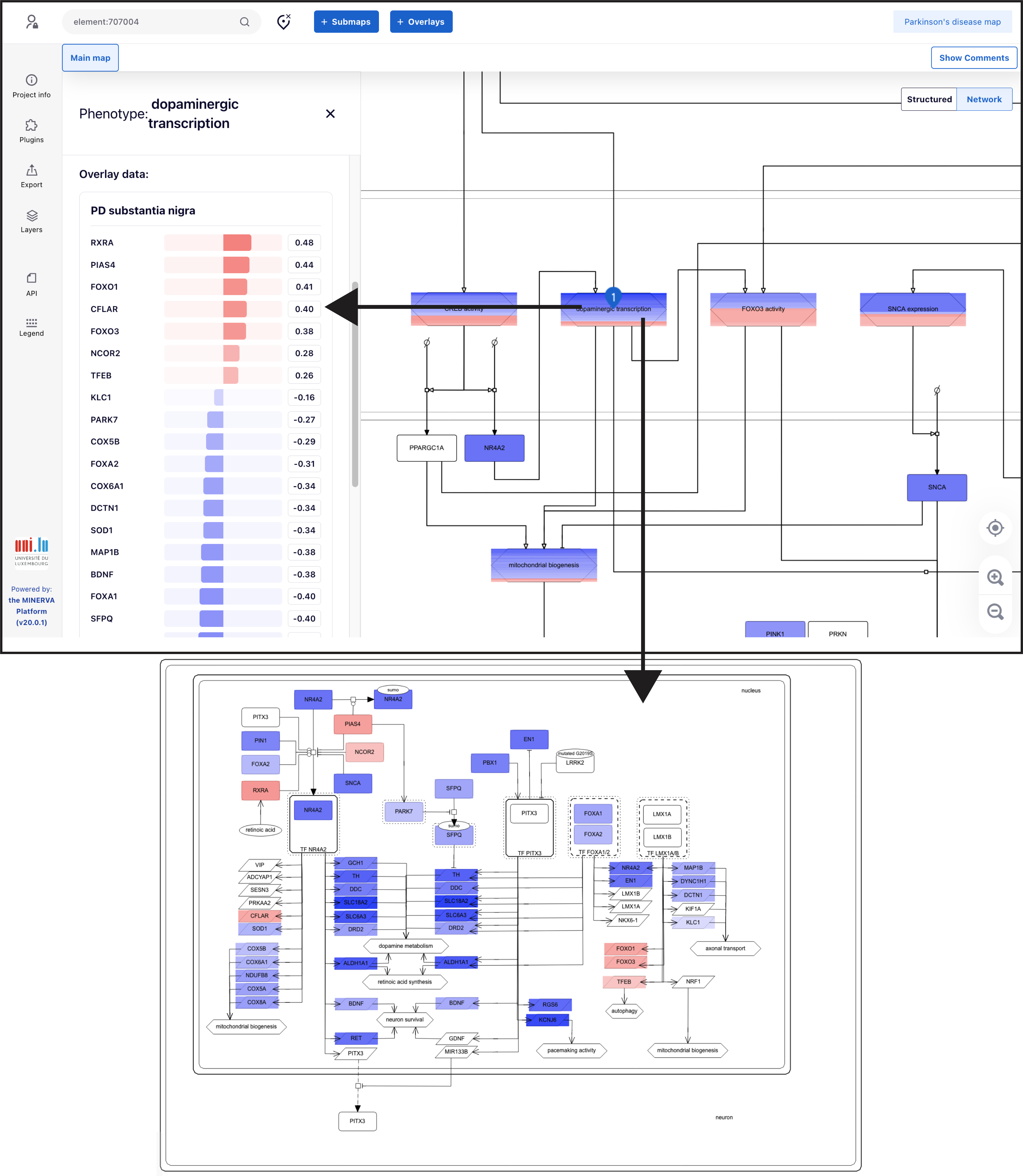
Overlays by value range#
Overlays can colour elements according to values from [-1,1] range. This allows visualisation of experimental data, for instance differentially expressed molecules. For that purpose, expression values have to be normalised to [-1,1] range, i.e., scaled so that values fall between -1 and 1. Visualised colour range can be configured individually for each MINERVA instance.
After clicking on an element of the map, overlay data for this element are visible in the left panel as one or more horizontal colour bars, together with the values.
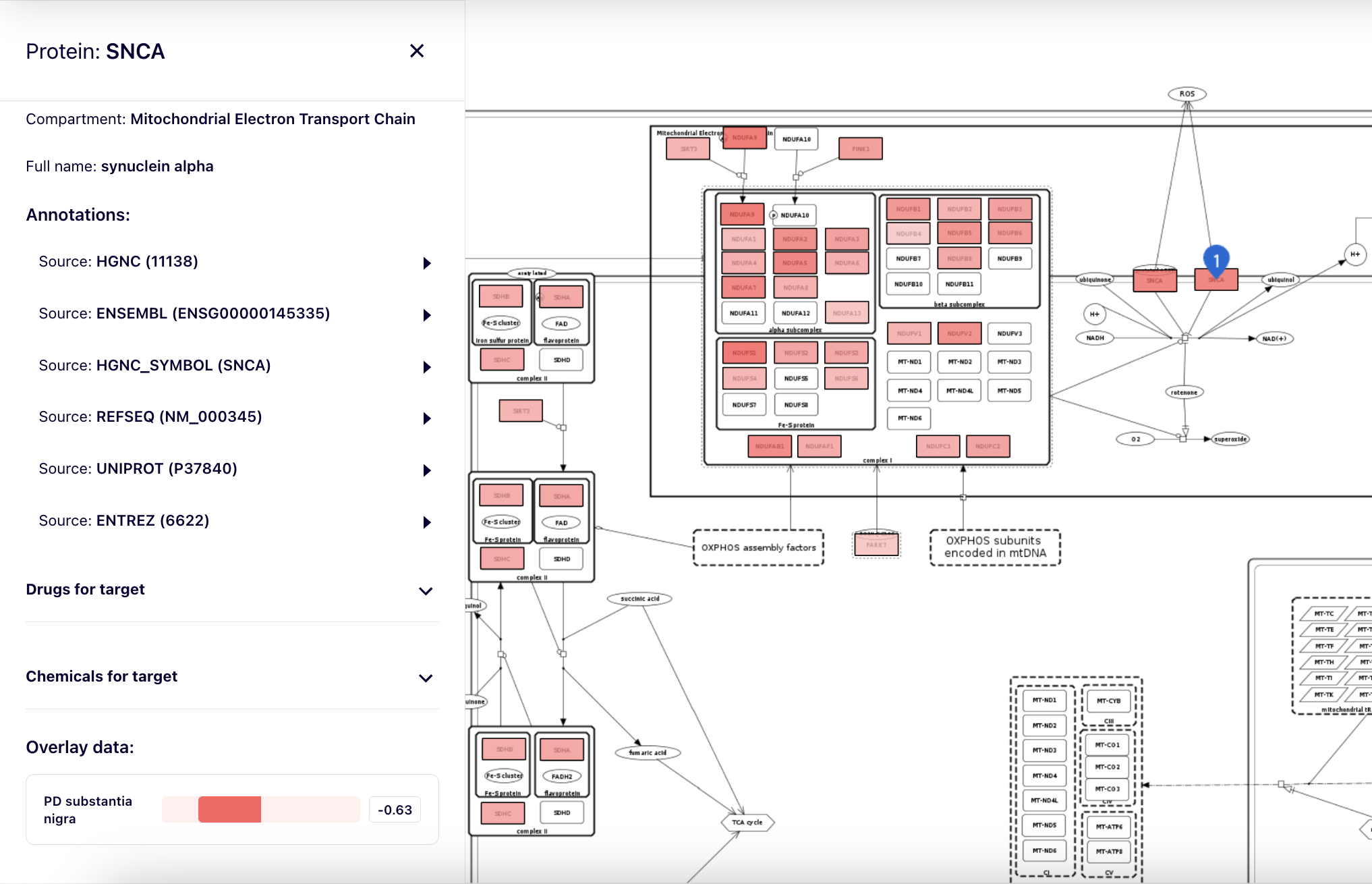
Overlays of genetic variants#
The genetic variants overlay is currently being redesigned and is not available in v20.0.
User provided Overlays#
The overlays provided by the user, are private and visible only to the currently logged-in user. Check all the data columns that can be used in an overlay file.
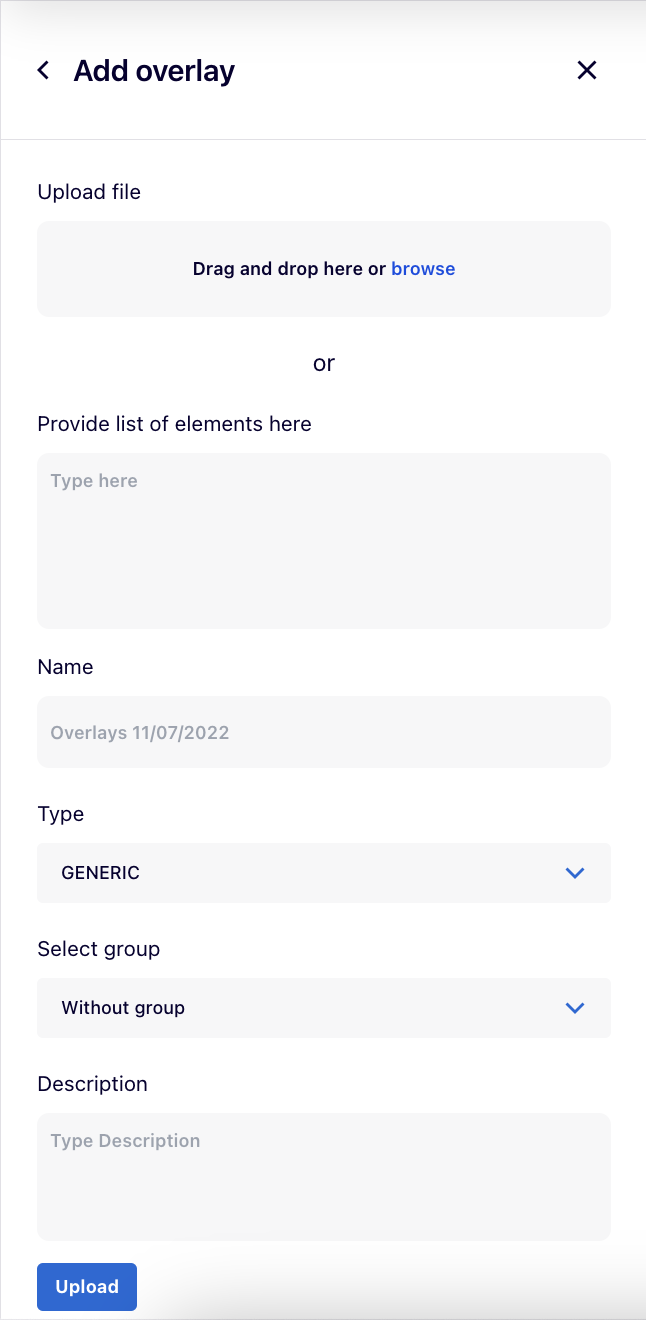
Using the “Add overlay” button, users can upload their own overlay:
- Upload file with the overlay’s dataset or Provide list of elements as the overlay’s dataset in the designated box.
- Name for the overlay.
- The Type field depends on the header of the uploaded file. After selecting it, the dropdown list for Type automatically switches between GENERIC and GENETIC_VARIANT, as MINERVA recognises the file’s header because genetic variant overlays require a specific header field.
- A GENERIC overlay displays colouring on the map according to the dataset of indicated molecules (e.g., differentially expressed genes).
- A GENETIC VARIANT overlay displays genetic variant data on the map. This overlay type is currently being redesigned and is not available in v20.0.
- Select group allows to choose a group to which an overlay will belong (see below).
- Description for the overlay (optional).
After clicking on “Upload” button, your overlay will become available in the left panel.
See overlay data examples for more information.
Overlay groups#
Users can organise their overlays into groups. “Add group” button allows to define a new group, which will then become available in the “Add overlay” menu (“Select group”). Existing overlays can be moved between the groups using the three dot button next to overlay names. Finally, group name and order can be changed using a similar button next to a group name.
Layers#
Layers provide an additional level of annotation on top of diagrams, highlighting specific areas or guiding users though the content.
They are accessed by using the Layers button in the left menu. Users can toggle the visibility of selected layers using the “eye” icon next to the layer name. The image below indicates, from left to right, the “Layers” button, “eye” button, and a component of a layer.
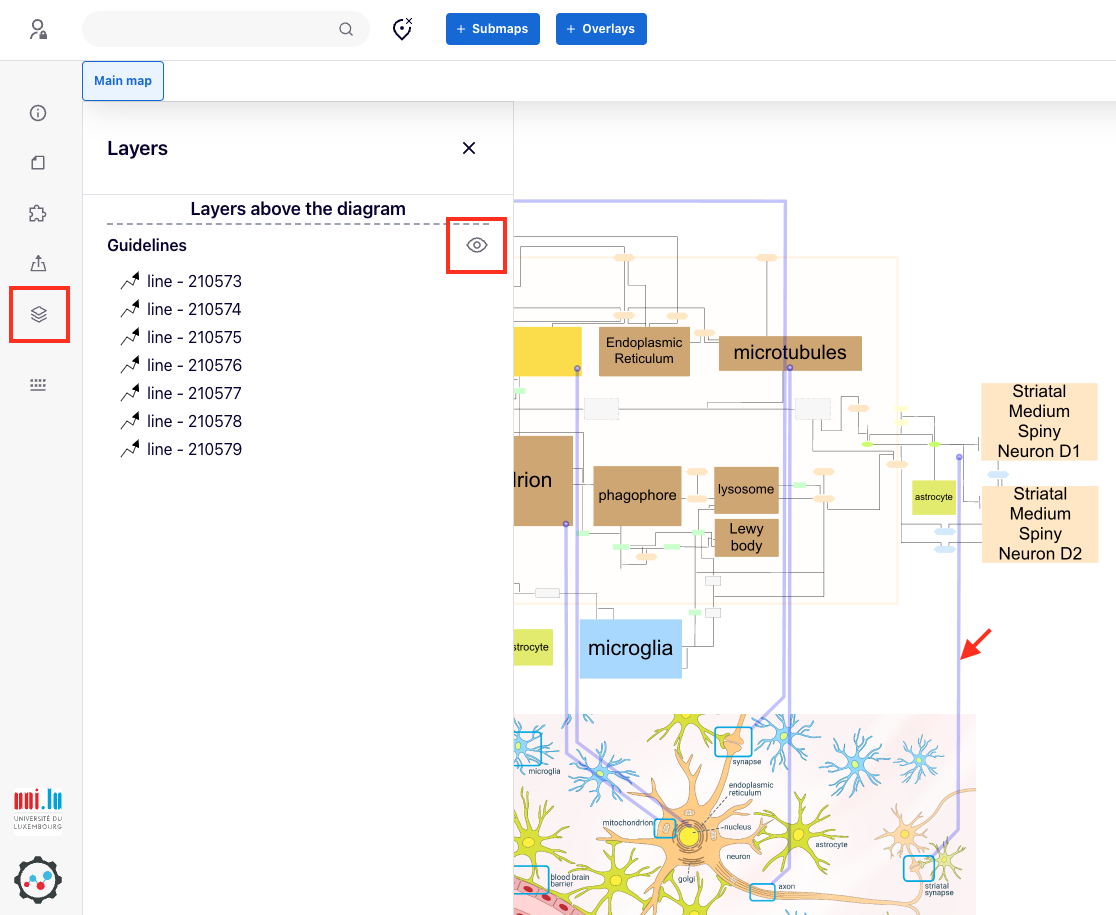
Layers can be constructed and edited by the administrators of a given MINERVA instance, or curators of a given map. See section layers in the administrator’s manual.
Plugins#
Plugins allow users to create custom visualisations in the browser, independent of the core functionalities of MINERVA.
By clicking on the ![]() icon, two panels, left and right, will appear.
icon, two panels, left and right, will appear.
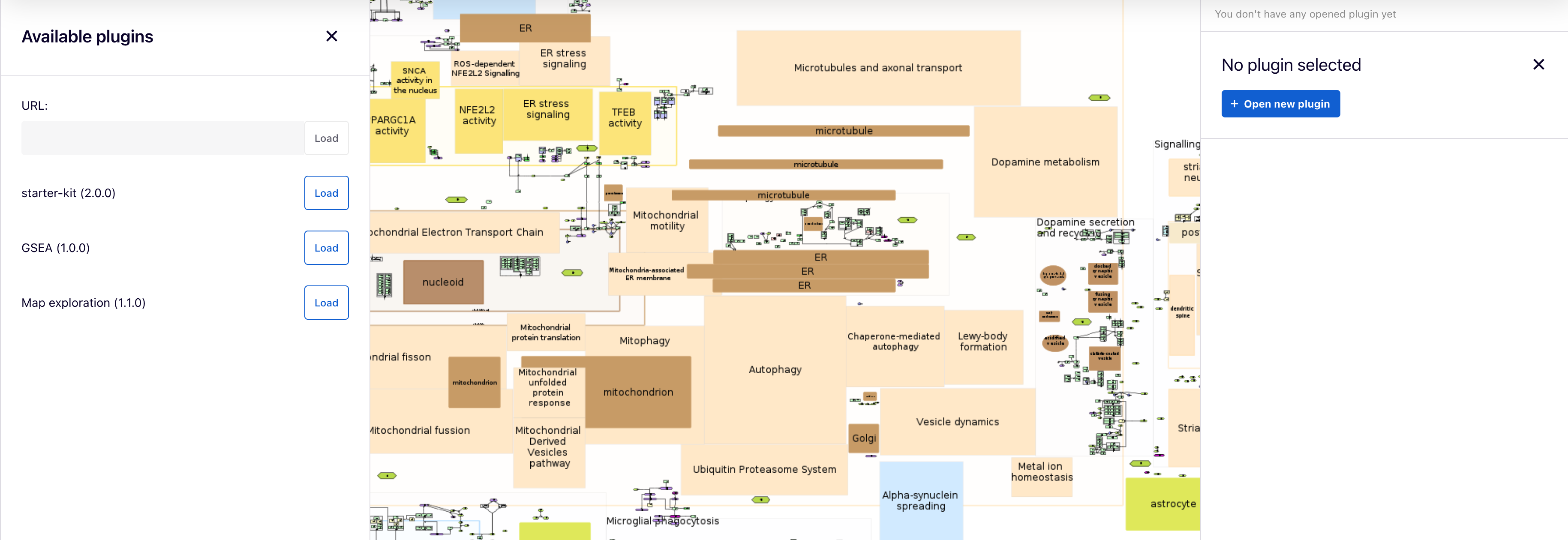
Available plugins for all users are uploaded by admins on the MINERVA platform and are listed in the left panel, where users can load each one. Also, each plugin can be selected in the right panel by clicking on the  button.
button.
Additionally, users can supply a URL to a plugin written in JavaScript in the URL field. To learn how to write your own plugins, visit the “Plugin starter kit” repository.
Example plugin: Enter the provided URL into the ‘URL’ field and click the ‘Load’ button:
https://minerva-service.lcsb.uni.lu/plugins/gsea/plugin.js
Please find below the list of available plugins, click at plugin’s name for its details.
| Name of the plugin | Address URL and the short description of the plugin |
|---|---|
| Map exploration | Enhance the visualisation and exploration of molecular interaction in the map. https://minerva-service.lcsb.uni.lu/plugins/exploration/plugin.js |
| GSEA | Calculate the enrichment of the elements from the selected overlay in the pathway. https://minerva-service.lcsb.uni.lu/plugins/gsea/plugin.js |