Back to the “Overlays” in user manual
Example 3 - Overlay data upload by registered users#
You will find here examples of data upload by registered users, for basic and advanced overlays.
Basic format#
The file in basic format has two tab-separated columns Name and Value (see also Upload user-provided overlay data):
- Name - names of elements to be coloured.
- Value - a value from [-1, 1] range, to be transformed to red-blue colouring, with negative values coloured red, positive coloured blue; the saturation of the colour will be defined by the absolute value.
Note Blue (for negative values) and red (for positive values) are configured in the admin panel (Overlays) and can differ between MINERVA instances.
Click here to download the basic overlay, containing a custom colouring for an example map.
To upload this dataset, do the following:
- Make sure you have the privileges to generate overlays in the example map.
- Go to Projects tab and Edit the example map.
- In the Users tab, check the box in the View project column.
- Open the example map from the Projects tab by clicking on its name (default: map_example).
- Make sure that you are logged in by checking the Info tab in the left panel.
- In Overlays > User-provided overlays section
- Choose Add overlay then press Choose File.
- Select basic overlay file to upload.
- In Overlays > User-provided overlays tab, click View checkbox to examine generated view.
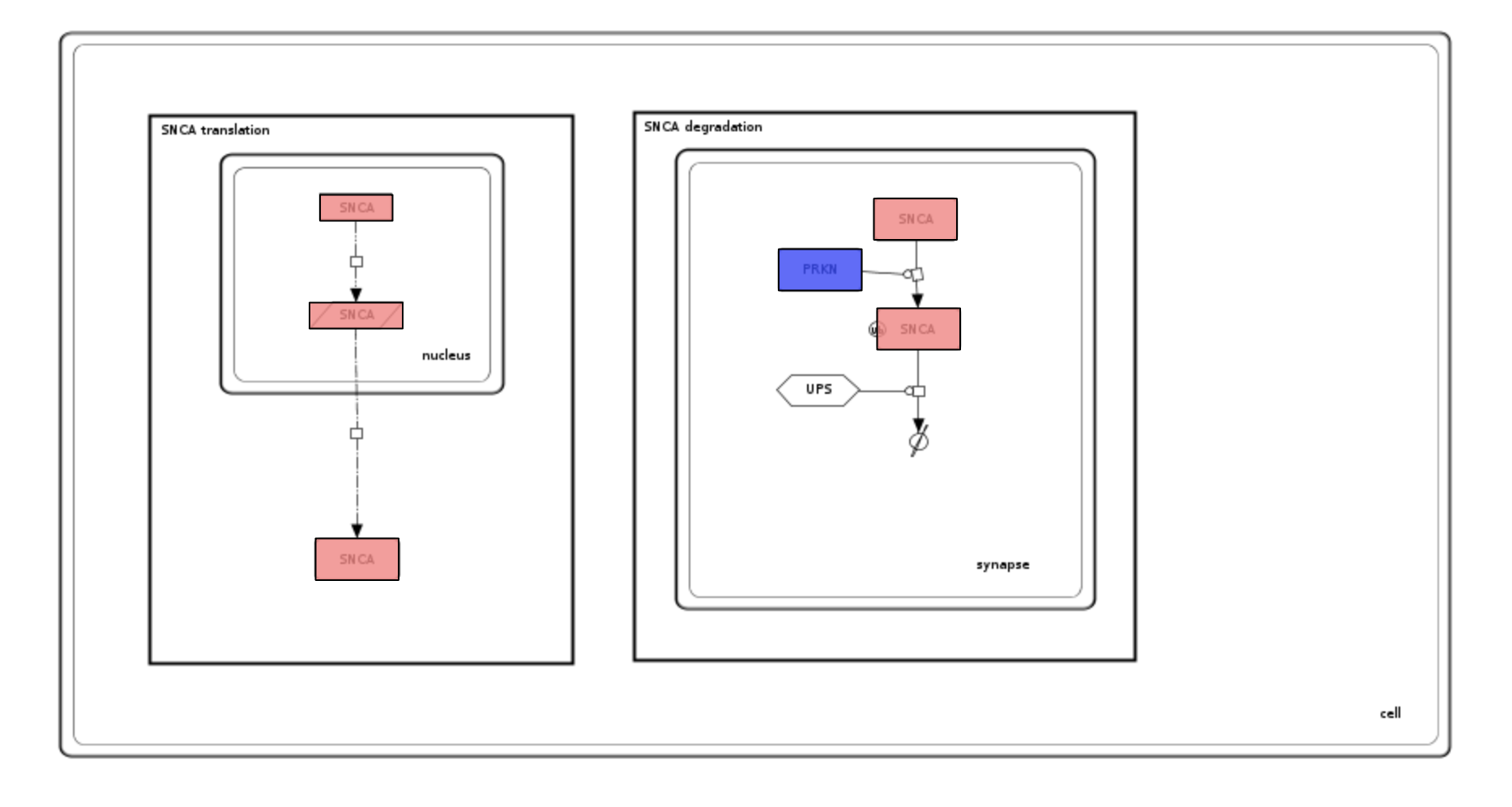
Examine the generated colouring and the uploaded dataset. Please see image below and note that:
- All SNCA elements are coloured to the same colour because of by-name matching.
- SNCA elements are red-coloured (the value in the file is -0.5).
- PRKN is blue (the value in the file is 0.75)
Advanced format#
Advanced format foresees two parts of the uploaded dataset - header and body (see also Upload user-provided overlay data).
The header lines start with #. It contains the following elements:
#VERSION=xyz- a version of this custom layout#NAME=xyz- a name that will be automatically assigned upon upload#DESCRIPTION=xyz- a description that will be automatically assigned upon upload
The body is a table with the following set of columns.
Warning! Colouring of the elements in different compartments works ONLY for separate (not nested) compartments. Setting a colour for elements in a compartment that encloses other compartments does not set the colour inside the nested compartments.
By-identifier colouring#
Click here to download the advanced overlay (generic), containing a custom colouring for an example map.
Upload this dataset as in basic format upload.
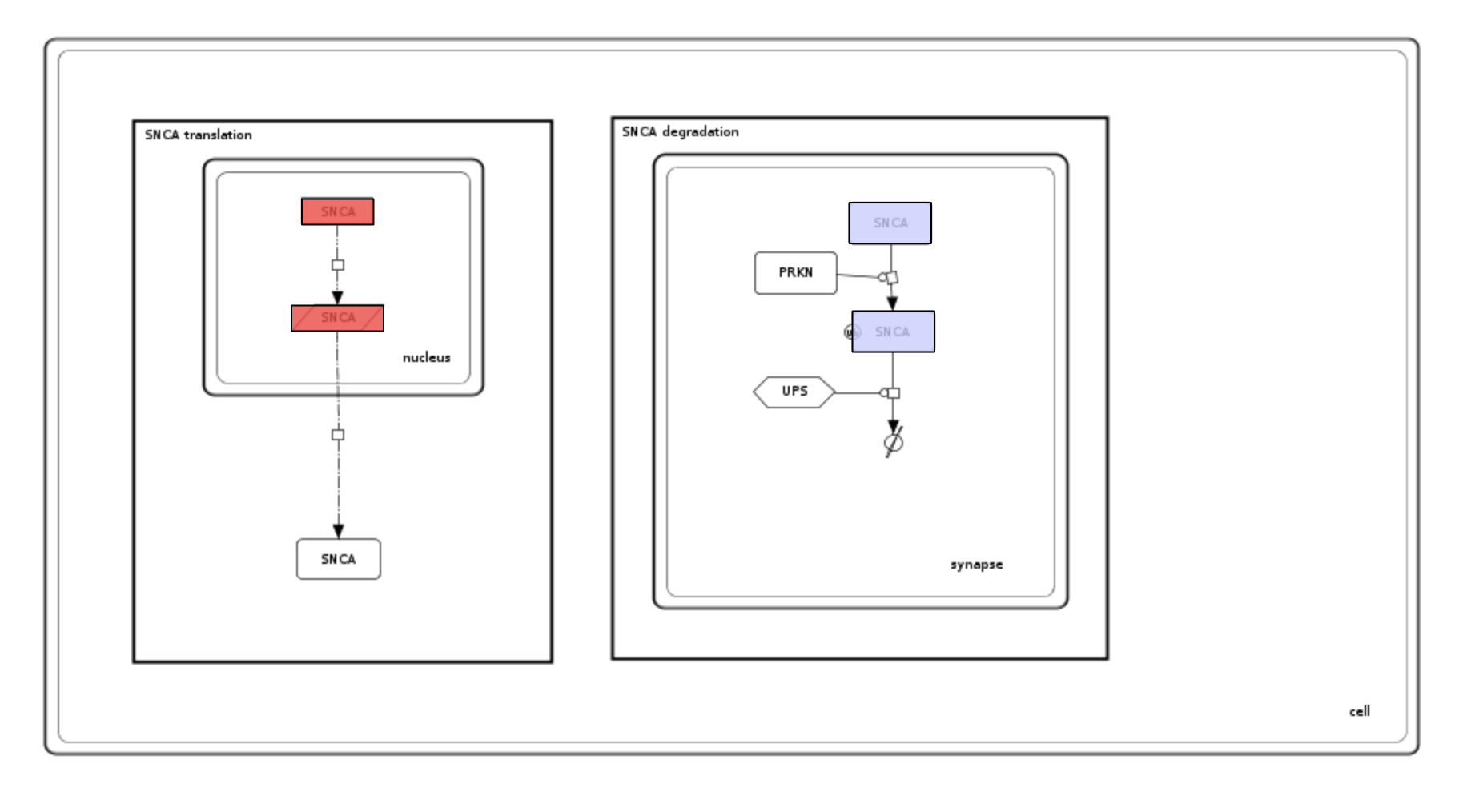
Examine the generated colouring and the uploaded dataset. Please see image below and note that:
- elements are coloured by their identifiers instead of their names,
- SNCA elements in the nucleus elements are coloured red, synapse elements are coloured blue due to by-compartment colouring.
- Phenotype UPS is not coloured as the smallest compartment UPS is contained in is synapse, not cell.
Warning! Colouring compartments works only for separate (not nested) compartments. For example: setting the colour for SNCA identifier in the cell compartment will not colour the contents of the nucleus and synapse.
Using SNCA colouring by-name overrides by-identifier colouring, the order of selection is as follows:
- match by-name
- match by-identifier
- constrain by-compartment
Custom colors and interaction colouring#
User-defined colors can be assigned to both elements and interactions, using the hex code (e.g. #fffaaa).
Click here to download the interactions overlay, containing a custom colouring for the example map.
Upload this dataset as described above in basic format upload.
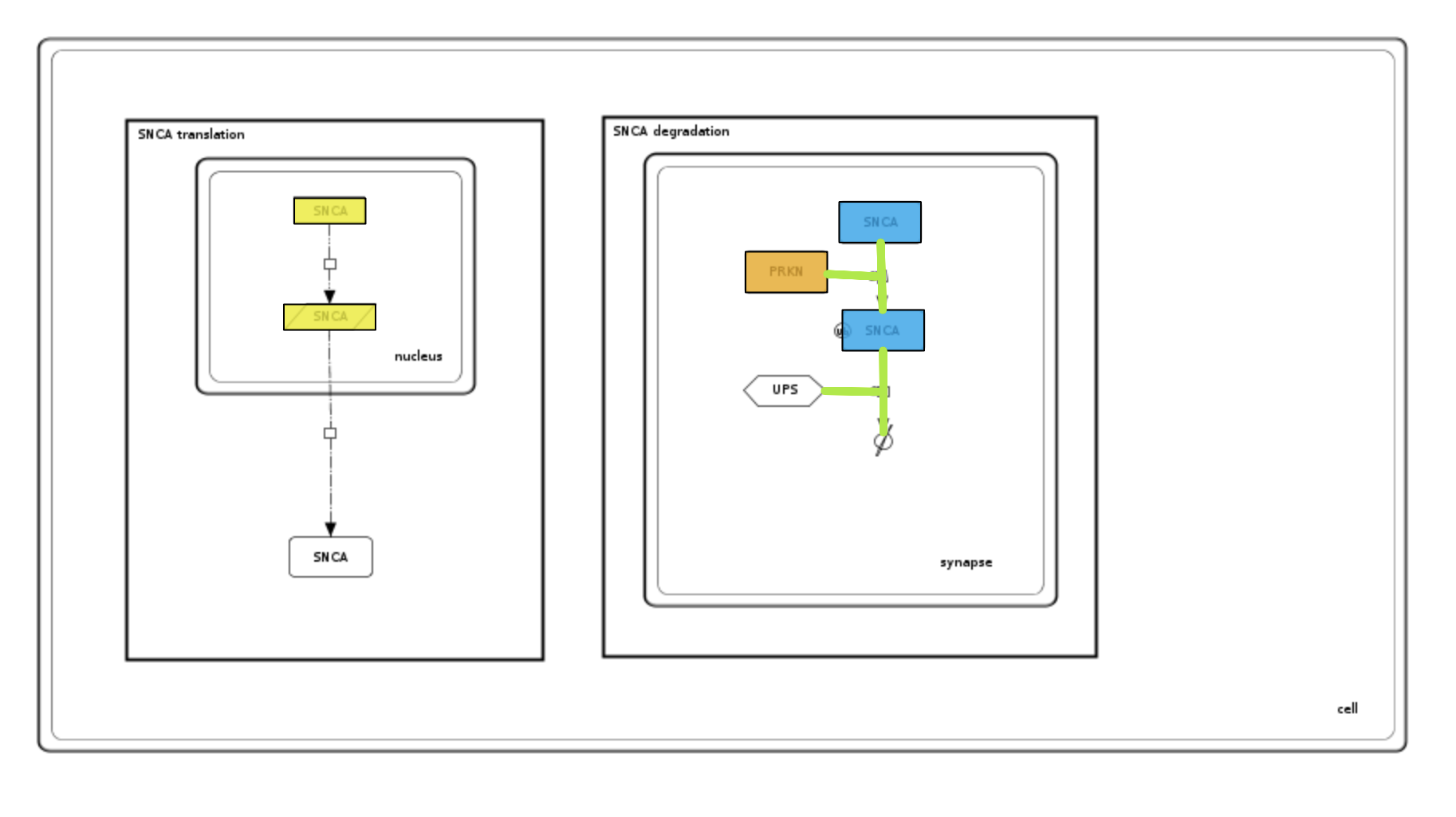
Examine the generated colouring and the uploaded dataset. Please see image below and note that:
- color column replaces value for defining colours of interactions and elements; both parameters cannot be used simultaneously.
- reaction id provided in element_identifier it is not constraint by compartment (as more than one compartment can contain reactions line).
Remarks#
- The uploaded dataset must be a text file.
- Column Name refers to the full name in the annotation panel; e.g., HGNC gene name for genes/proteins; ChEBI name for simple molecules or GO name for a Gene Ontology term.
- Identifiers of the elements in the dataset have to follow the MIRIAM standard; e.g., HGNC: 11138; Ensembl: ENSG00000184845; ChEBI: CHEBI:18243.
- Most columns can be used simultaneously. If the same element is addressed in two different columns, it will be coloured with these two colors.
- Restrictions:
- Either a value or a color columns is necessary in the uploaded dataset, but they mutually exclusive.
- Reactions to be coloured should be identified only by reaction id or PubMed.
- If the compartment column is used, it is highly recommended to fill all its fields with a compartments name from the map. Element which does not have defined compartment will be coloured in the whole map.
Alternatively You can also export a diagram of interest as a CellDesigner file, open it in CellDesigner and colour it there.